 ++++
++++Notebook converted from Jupyter for blog publishing.
00-AdaBoost
AdaBoost
The Data

Mushroom Hunting: Edible or Poisonous?
Data Source: https://archive.ics.uci.edu/ml/datasets/Mushroom (opens in a new tab)
This data set includes descriptions of hypothetical samples corresponding to 23 species of gilled mushrooms in the Agaricus and Lepiota Family (pp. 500-525). Each species is identified as definitely edible, definitely poisonous, or of unknown edibility and not recommended. This latter class was combined with the poisonous one. The Guide clearly states that there is no simple rule for determining the edibility of a mushroom; no rule like ``leaflets three, let it be'' for Poisonous Oak and Ivy.
Attribute Information:
- cap-shape: bell=b,conical=c,convex=x,flat=f, knobbed=k,sunken=s
- cap-surface: fibrous=f,grooves=g,scaly=y,smooth=s
- cap-color: brown=n,buff=b,cinnamon=c,gray=g,green=r, pink=p,purple=u,red=e,white=w,yellow=y
- bruises?: bruises=t,no=f
- odor: almond=a,anise=l,creosote=c,fishy=y,foul=f, musty=m,none=n,pungent=p,spicy=s
- gill-attachment: attached=a,descending=d,free=f,notched=n
- gill-spacing: close=c,crowded=w,distant=d
- gill-size: broad=b,narrow=n
- gill-color: black=k,brown=n,buff=b,chocolate=h,gray=g, green=r,orange=o,pink=p,purple=u,red=e, white=w,yellow=y
- stalk-shape: enlarging=e,tapering=t
- stalk-root: bulbous=b,club=c,cup=u,equal=e, rhizomorphs=z,rooted=r,missing=?
- stalk-surface-above-ring: fibrous=f,scaly=y,silky=k,smooth=s
- stalk-surface-below-ring: fibrous=f,scaly=y,silky=k,smooth=s
- stalk-color-above-ring: brown=n,buff=b,cinnamon=c,gray=g,orange=o, pink=p,red=e,white=w,yellow=y
- stalk-color-below-ring: brown=n,buff=b,cinnamon=c,gray=g,orange=o, pink=p,red=e,white=w,yellow=y
- veil-type: partial=p,universal=u
- veil-color: brown=n,orange=o,white=w,yellow=y
- ring-number: none=n,one=o,two=t
- ring-type: cobwebby=c,evanescent=e,flaring=f,large=l, none=n,pendant=p,sheathing=s,zone=z
- spore-print-color: black=k,brown=n,buff=b,chocolate=h,green=r, orange=o,purple=u,white=w,yellow=y
- population: abundant=a,clustered=c,numerous=n, scattered=s,several=v,solitary=y
- habitat: grasses=g,leaves=l,meadows=m,paths=p, urban=u,waste=w,woods=d
Goal
THIS IS IMPORTANT, THIS IS NOT OUR TYPICAL PREDICTIVE MODEL!
Our general goal here is to see if we can harness the power of machine learning and boosting to help create not just a predictive model, but a general guideline for features people should look out for when picking mushrooms.
Imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as snsdf = pd.read_csv("../DATA/mushrooms.csv")df.head()class
cap-shape
cap-surface
cap-color
bruisesEDA
sns.countplot(data=df,x='class')<AxesSubplot:xlabel='class', ylabel='count'>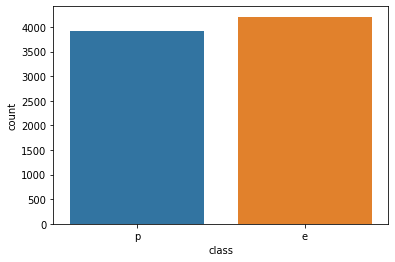
df.describe()class
cap-shape
cap-surface
cap-color
bruisesdf.describe().transpose()count
unique
top
freq
classplt.figure(figsize=(14,6),dpi=200)
sns.barplot(data=df.describe().transpose().reset_index().sort_values('unique'),x='index',y='unique')
plt.xticks(rotation=90);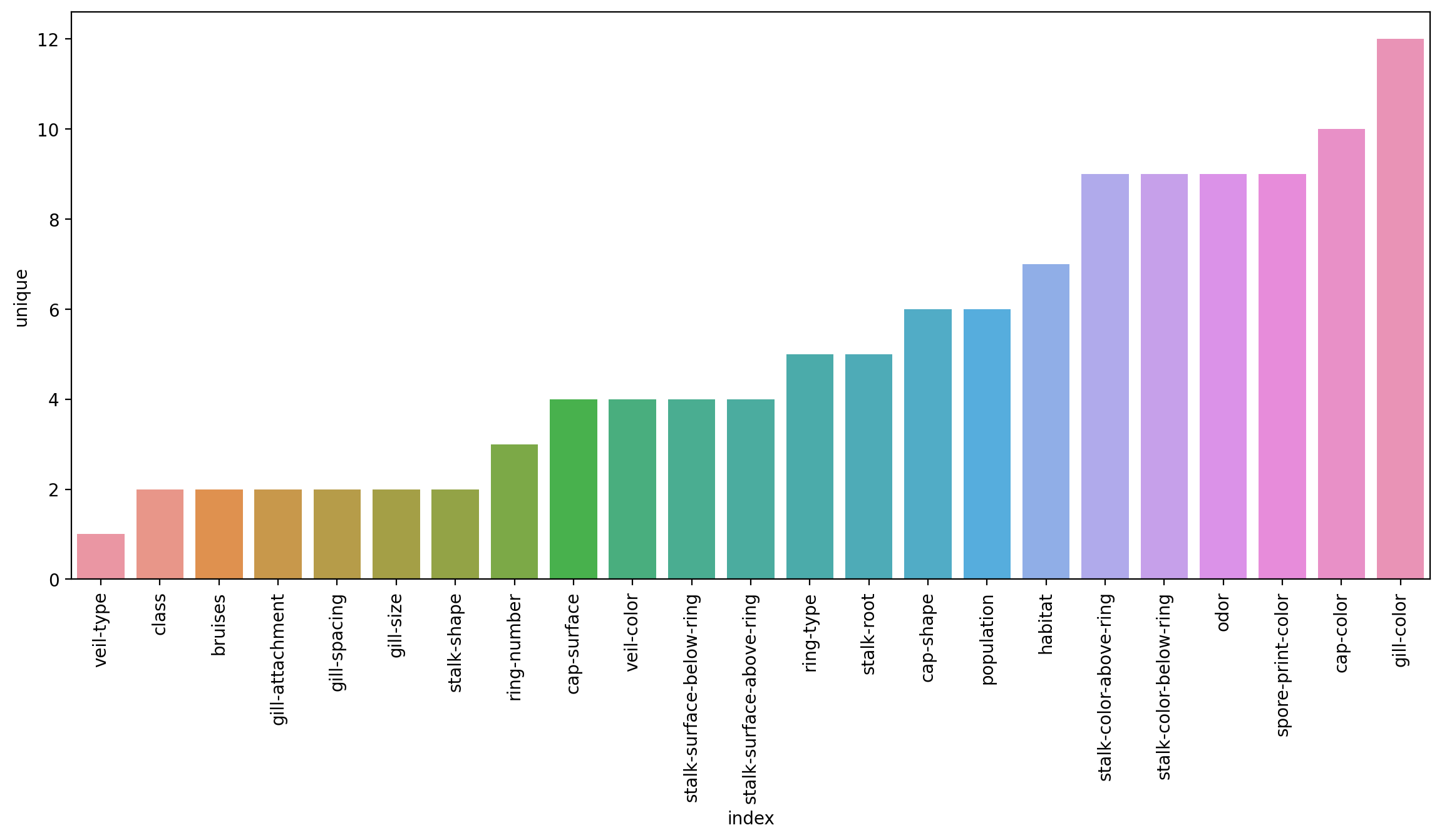
Train Test Split
X = df.drop('class',axis=1)X = pd.get_dummies(X,drop_first=True)y = df['class']from sklearn.model_selection import train_test_splitX_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.15, random_state=101)Modeling
from sklearn.ensemble import AdaBoostClassifiermodel = AdaBoostClassifier(n_estimators=1)model.fit(X_train,y_train)AdaBoostClassifier(n_estimators=1)Evaluation
from sklearn.metrics import classification_report,plot_confusion_matrix,accuracy_scorepredictions = model.predict(X_test)predictionsarray(['p', 'e', 'p', ..., 'p', 'p', 'e'], dtype=object)print(classification_report(y_test,predictions)) precision recall f1-score support
e 0.96 0.81 0.88 655
p 0.81 0.96 0.88 564
model.feature_importances_array([0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.,
0., 0., 0., 0., 0., 1., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.,
0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.,
0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.,
0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.,model.feature_importances_.argmax()22X.columns[22]'odor_n'sns.countplot(data=df,x='odor',hue='class')<AxesSubplot:xlabel='odor', ylabel='count'>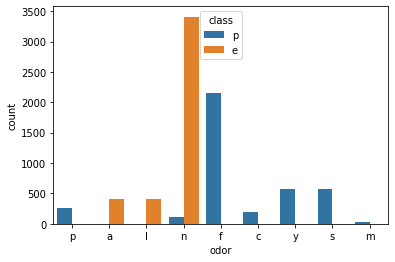
Analyzing performance as more weak learners are added.
len(X.columns)95error_rates = []
for n in range(1,96):
model = AdaBoostClassifier(n_estimators=n)
model.fit(X_train,y_train)
preds = model.predict(X_test)
err = 1 - accuracy_score(y_test,preds)
error_rates.append(err)plt.plot(range(1,96),error_rates)[<matplotlib.lines.Line2D at 0x289c33b1f70>]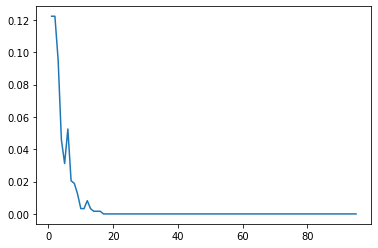
modelAdaBoostClassifier(n_estimators=95)model.feature_importances_array([0. , 0. , 0. , 0. , 0. ,
0. , 0. , 0. , 0.01052632, 0. ,
0. , 0.01052632, 0. , 0. , 0. ,
0.01052632, 0. , 0.05263158, 0.03157895, 0.03157895,
0. , 0. , 0.06315789, 0.02105263, 0. ,feats = pd.DataFrame(index=X.columns,data=model.feature_importances_,columns=['Importance'])featsImportance
cap-shape_c
0.000000
cap-shape_f
0.000000imp_feats = feats[feats['Importance']>0]imp_featsImportance
cap-color_c
0.010526
cap-color_n
0.010526imp_feats = imp_feats.sort_values("Importance")plt.figure(figsize=(14,6),dpi=200)
sns.barplot(data=imp_feats.sort_values('Importance'),x=imp_feats.sort_values('Importance').index,y='Importance')
plt.xticks(rotation=90);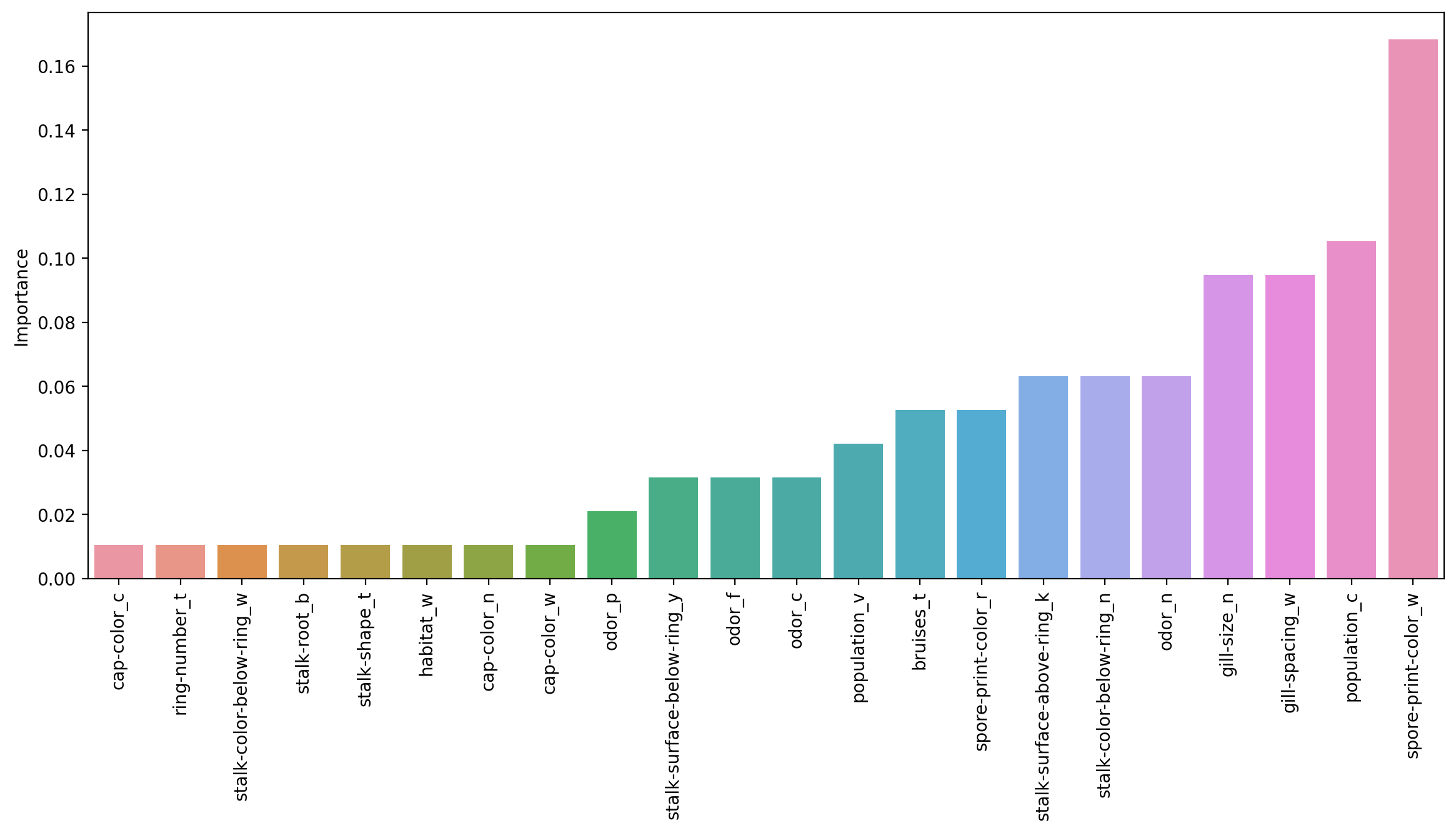
sns.countplot(data=df,x='habitat',hue='class')<AxesSubplot:xlabel='habitat', ylabel='count'>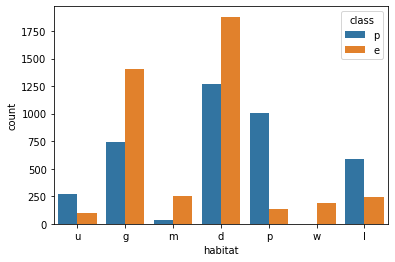
Interesting to see how the importance of the features shift as more are allowed to be added in! But remember these are all weak learner stumps, and feature importance is available for all the tree methods!